What is a variant?
The human genome contains over 3 billion base pairs– individual “letters” in the DNA code that make the instructions for our cells and bodies to function. A variant or mutation is any change in this code. These changes can be passed down from our biological parents or arise independently. Each variant can range from a single “letter” to massive stretches of DNA. Even though we share 99.9% of our genome with each other, we each still vary at 3 million base pairs!
While some variants or mutations in our genes cause disease, many do not. Each variant is placed in a category on a spectrum based on how confident we can be that it is benign or pathogenic. Benign variants do not contribute to a disease or condition, whereas pathogenic variants contribute to or cause a disease or condition.
What is a VUS?
When we do not have enough information to know whether a variant is or is not associated with a given condition, the variant is considered a variant of uncertain significance (VUS). Importantly, just because a variant is new does not mean it is a VUS. Instead, certain rules are used to sort variants into categories along the spectrum of VUS, benign, or pathogenic. It is important to note that these categories reflect how confident we can be that the variant causes or does not cause a condition, not how strong its effect on our health may be.
After a VUS is identified, more information may allow healthcare professionals and researchers to reclassify the variant along the spectrum. This happens when we become more confident that a variant is either benign or pathogenic. We can think of this like a balance scale (Figure 1). As more evidence about a variant accumulates on one arm, the scale leans towards that side. For a VUS, the scale is balanced. Research methods used to (re)classify a variant range from computer modeling to laboratory studies to searching the population for a variant. As more patients are identified with the same variant, patterns emerge that help researchers understand whether the variant is likely to be disease-causing. For example, if many people without a given condition have the same VUS, it is less likely to be pathogenic. Conversely, if a variant appears repeatedly in people with the condition but not in unaffected individuals, this suggests it may be pathogenic. Over time, as evidence accumulates, the classification can shift from VUS to either benign or pathogenic. In general, it is more common for a VUS to be reclassified as a benign variant than a pathogenic one.
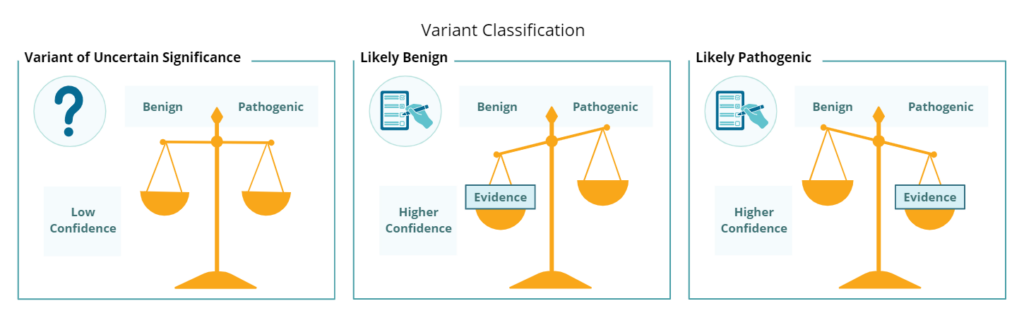
Figure 1: Variant Classification. Variants in our genome are classified based on the weight of evidence available to us. Like a balance scale, as more evidence accumulates, we can be more confident when considering how a variant might affect our health. In the left panel, the lack of evidence characterizes a VUS. In the middle panel, the accumulation of evidence suggests a variant is likely benign. In the right panel, the accumulation of evidence suggests a variant is likely pathogenic. Created in BioRender. Walker, K. (2025) https://BioRender.com/qm0yuic.
How is a VUS identified?
Any one of us may receive genetic testing, or DNA sequencing, for a number of reasons. If you would like to learn more about genetic testing for Ataxias, please take a look at the Genetic Testing page, or this article for a brief overview. When we provide a sample of DNA for testing, the sample is sent to a genetic testing laboratory where a series of machines using chemical processes reads each letter in the DNA code. By comparing the unique code with known information about variants and genes, the laboratory creates a genetic testing report. If you have your genome or one or more of your genes sequenced, your genetic testing report may tell you and your care team that you carry a VUS.
A VUS is not considered clinically actionable, meaning it would not be used to make healthcare decisions at the time it is identified. Your care team will look at the report and consider your individual symptoms and history to interpret this information and work with you to make a plan for your care. If you or a family member receives a VUS result, it is important to discuss this with a genetic counselor, who is specifically trained to help patients understand what a VUS means for their specific situation. Genetic counselors can also advise whether periodic re-evaluation of the variant’s classification is recommended, as variants that are uncertain today may be better understood in the future as more evidence accumulates.
If you would like to learn more about Variant of Uncertain Significance, take a look at these resources by The NHS and Genomics England.
Written by Keenan Walker and edited by Larissa Nitschke, PhD.
Read Other SCAsource Summary Articles

Snapshot: What is a CT Scan?
A computer tomography (CT) scan, also called a CAT scan, is a diagnostic imaging procedure that uses X-rays and a computer to help doctors see inside your body. CT scans Read More…
An unexpected guest found in toxic ataxin-1 clumps in SCA1
Written by Anastasiya Potapenko Edited by Priscila Pereira Sena New clues into SCA1: RNA gets trapped inside toxic ataxin-1 clumps in brain cells, disrupting the production of proteins and contributing Read More…

Corriendo con la energía en vacío: Comprendiendo como la fatiga afecta la calidad de vida en las ataxias espinocerebelosas
Escrito por Alexandra Putka Editado por la Dra. Pragya Goel Traducido por Daphne Rincon ¿Te sientes cansado(a)? No estás solo(a). La fatiga es un síntoma común de las ataxias espinocerebelosas y afecta la calidad de vida. Las ataxias espinocerebelosas Read More…



